Part of series:Visualize
QQ图的各种画法
各种qq图的画法,简单示例
CMplot: Single trait
CMplot(pig60K,plot.type="q",box=FALSE,file="jpg",file.name=NULL,dpi=300,
conf.int=TRUE,conf.int.col=NULL,threshold.col="red",threshold.lty=2,
file.output=TRUE,verbose=TRUE,width=5,height=5)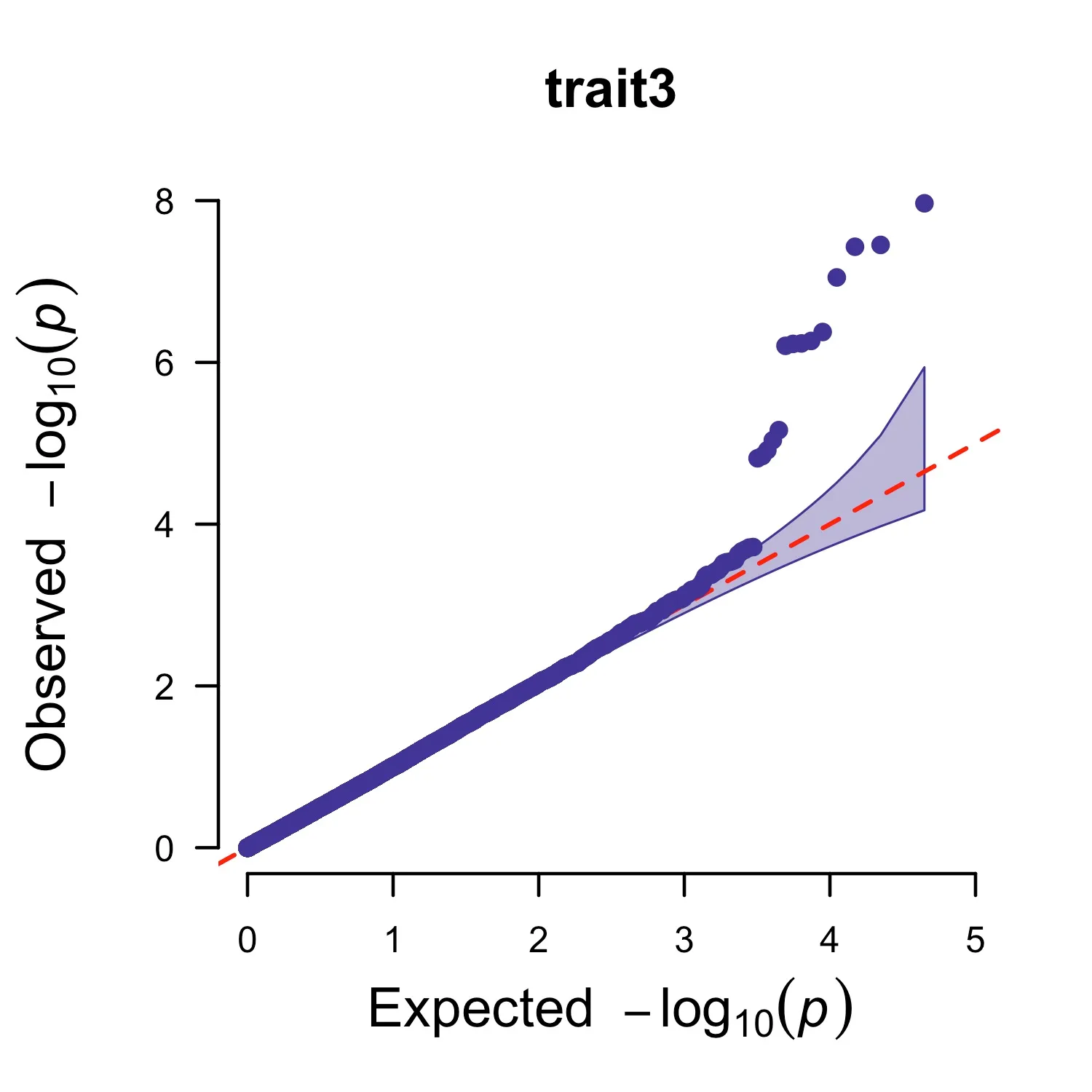
CMplot: Multi traits
CMplot(pig60K,plot.type="q",col=c("dodgerblue1", "olivedrab3", "darkgoldenrod1"),multraits=TRUE,
threshold=1e-6,ylab.pos=2,signal.pch=c(19,6,4),signal.cex=1.2,signal.col="red",
conf.int=TRUE,box=FALSE,axis.cex=1,file="jpg",file.name=NULL,dpi=300,file.output=TRUE,
verbose=TRUE,ylim=c(0,8),width=5,height=5)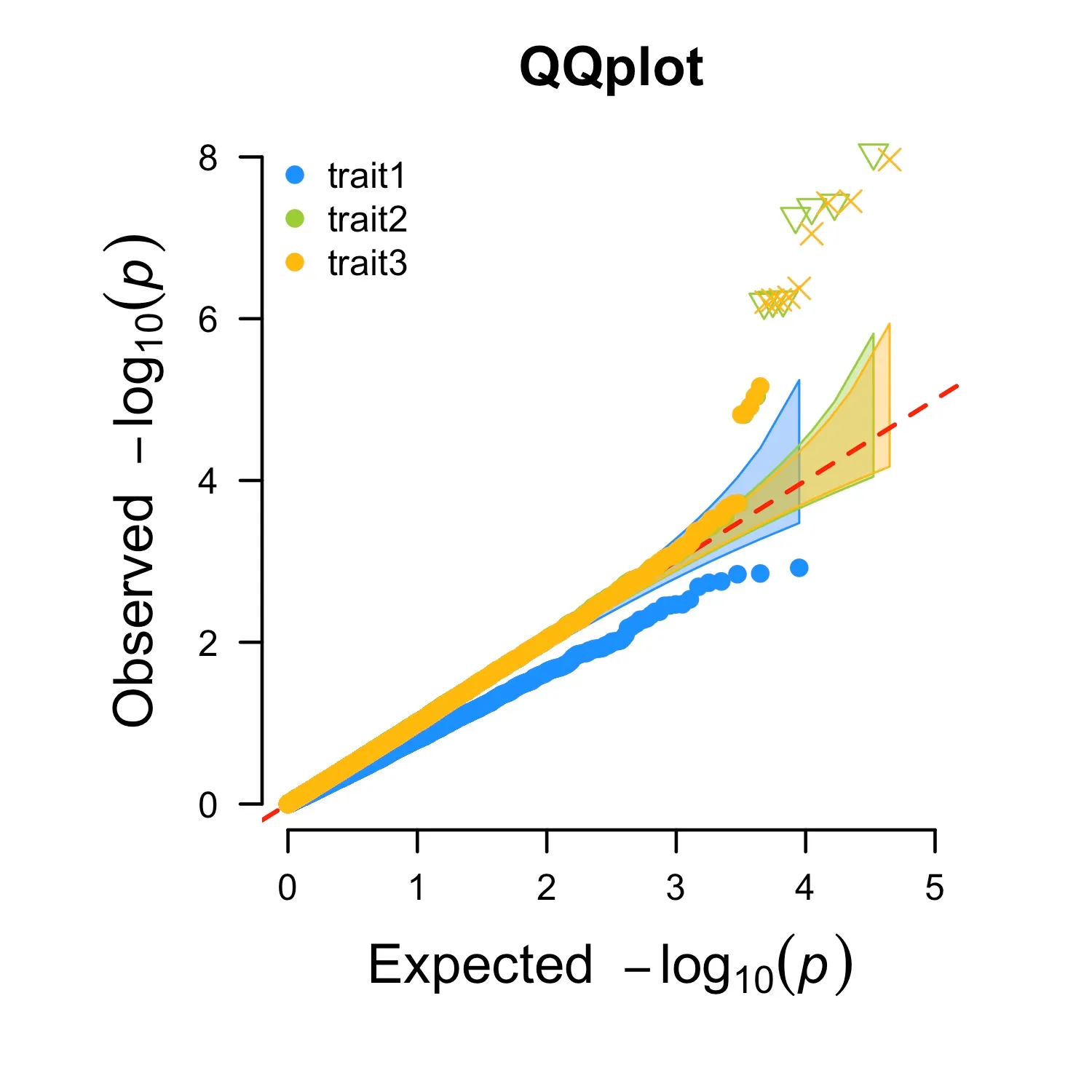
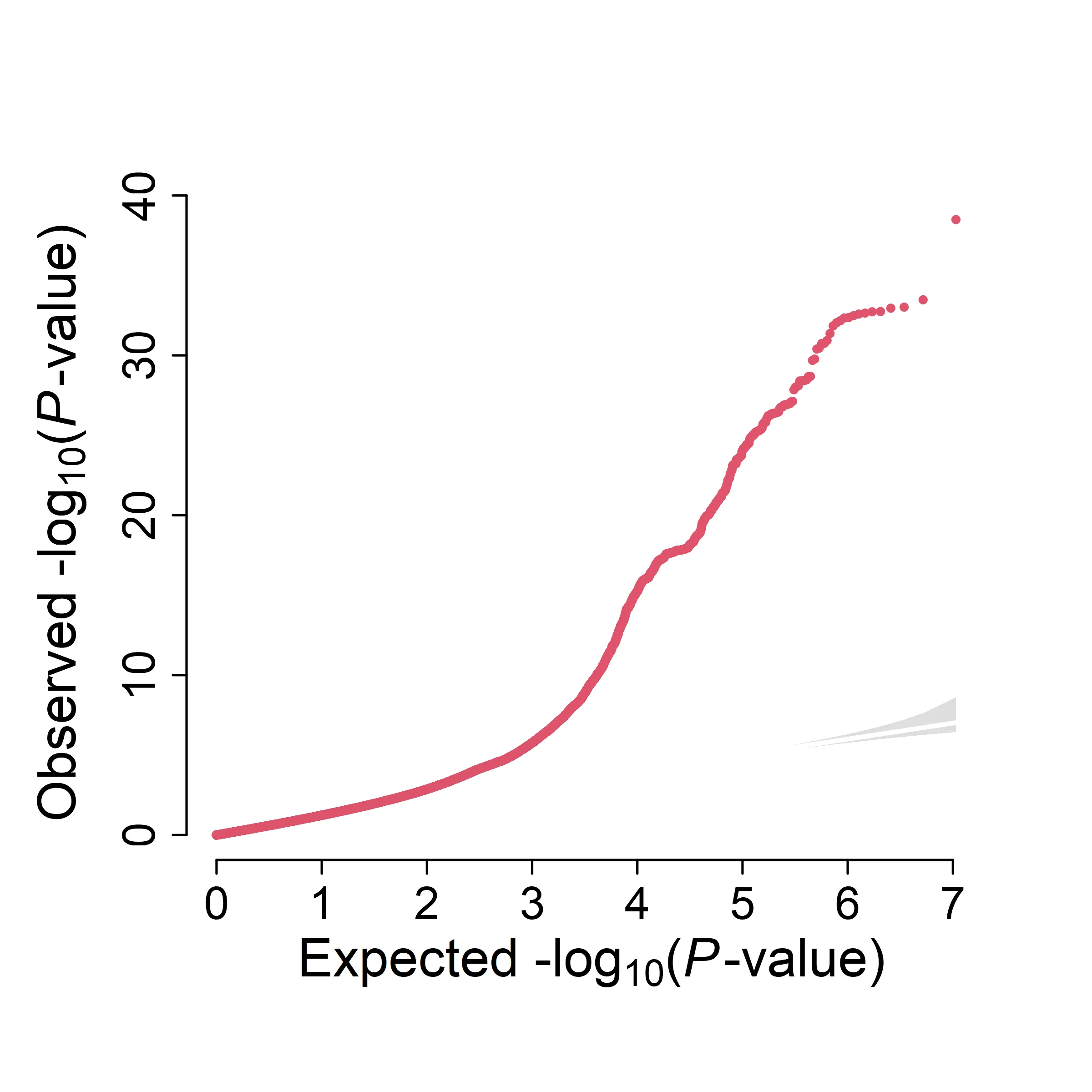
Custom Script: single trait
qqplot = function(pval, ylim=0, lab=1.4, axis=1.2){
par(mgp=c(5,1,0))
p1 <- pval
p2 <- sort(p1)
n <- length(p2)
k <- c(1:n)
alpha <- 0.05
lower <- qbeta(alpha/2, k, n+1-k)
upper <- qbeta((1-alpha/2), k, n+1-k)
expect <- (k-0.05)/n
biggest <- ceiling(max(-log10(p2), -log10(expect)))
xlim <- max (-log10(expect)+0.1);
if (ylim==0) ylim=biggest;
plot(-log10(expect), -log10(p2), xlim=c(0, xlim), ylim=c(0, ylim),
ylab=expression(paste("Observed ", "-", log[10], "(", italic(P), "-value)", sep="")),
xlab=expression(paste("Expected ", "-", log[10], "(", italic(P), "-value)", sep="")),
type="n", mgp=c(2,0.5,0), tcl=-0.3, bty="n", cex.lab=lab, cex.axis=axis)
polygon(c(-log10(expect), rev(-log10(expect))), c(-log10(upper), rev(-log10(lower))),
col=adjustcolor("grey", alpha.f=0.5), border=NA)
abline(0,1,col="white", lwd=2)
points(-log10(expect), -log10(p2), pch=20, cex=0.6, col=2)
}
Custom Script: multi trait
qqplot_multi <- function(pval_list, ylim=0, lab=1.4, axis=1.2, p_floor=1e-300, cols=NULL, pch=20, cex=0.6,
legend=TRUE, legend_pos="topleft", legend_cex=0.9, show_ci=TRUE, ci_scale=1, ci_alpha=0.6, ci_border=TRUE, diag_col="white", ...) {
if (!is.list(pval_list)) pval_list=as.list(as.data.frame(pval_list))
trait = names(pval_list)
if(is.null(trait) || any(trait == "")){
trait=paste0("Trait", seq_along(pval_list))
names(pval_list)=trait
}
m = length(pval_list)
if(m < 1) stop("pval_list must contain at least one trait ! ! !")
recycle1m = function(x, nm, name){
if (length(x) == 1) return(rep(x, nm))
if (length(x) == nm) return(x)
stop(name, " length must be 1 or ", nm, ".")
}
ci_scale = recycle1m(ci_scale , m, "ci_xfrac")
ci_scale[ci_scale <= 0] = 0.01
ci_scale[ci_scale > 1] = 1
if(is.null(cols)){
if ("hcl.colors" %in% getNamespaceExports("grDevices")){
cols = grDevices::hcl.colors(m, palette="Dark 3")
} else {
cols = grDevices::rainbow(m)
}
}
cols = recycle1m(cols, m, "cols")
pch = recycle1m(pch, m, "pch")
lighten_col = function(col, amount=0.65, alpha=ci_alpha) {
rgb = grDevices::col2rgb(col) / 255
rgb2 = rgb + (1-rgb) * amount
grDevices::rgb(rgb2[1,], rgb2[2,], rgb2[3,], alpha=alpha)
}
dat=vector("list", m)
for(i in seq_len(m)){
p = pval_list[[i]]
p = p[is.finite(p) & !is.na(p)]
p = p[p <= 1 & p >= 0]
if (length(p) == 0) stop("Trait '", trait[i], "' has no valid P in [0,1].")
p[p == 0] = p_floor
p2 = sort(p)
n = length(p2)
k = seq_len(n)
alpha = 0.05
lower = qbeta(alpha/2, k, n+1-k)
upper = qbeta(1 - alpha/2, k, n+1-k)
expect = (k-0.05) / n
expect[expect <= 0] = min(expect[expect > 0])
dat[[i]] = list(x=-log10(expect), y=-log10(p2), y_low=-log10(upper), y_high=-log10(lower), n=n)
}
xlim = max(vapply(dat, function(d) max(d$x, na.rm=TRUE), numeric(1))) + 0.1
ymax = max(vapply(dat, function(d) max(d$y, d$y_high, na.rm=TRUE), numeric(1)))
if (ylim==0) ylim=ceiling(ymax)
par(mgp=c(5,1,0))
plot(NA, xlim=c(0, xlim), ylim=c(0, ylim),
ylab=expression(paste("Observed", " -", log[10], "(", italic(P), "-value", ")", sep="")),
xlab=expression(paste("Expected", " -", log[10], "(", italic(P), "-value", ")", sep="")),
type="n", mgp=c(2,0.5,0), tcl=-0.3, bty="n", cex.lab=lab, cex.axis=axis, ...)
if (show_ci){
for(i in seq_len(m)){
d = dat[[i]]
o = order(d$x)
x = d$x[o]
ylow = d$y_low[o]
yhigh = d$y_high[o]
x_ci = x * ci_scale[i]
ylow = ylow * ci_scale[i]
yhigh = yhigh * ci_scale[i]
ci_col = lighten_col(cols[i])
if (ci_border) {
border=cols[i]
} else {
border=NA
}
polygon(c(x_ci, rev(x_ci)), c(ylow, rev(yhigh)), col=ci_col, border=border)
}
}
abline(0, 1, col=diag_col, lwd=2)
for(i in seq_len(m)){
d = dat[[i]]
points(d$x, d$y, pch=pch[i], cex=cex, col=cols[i])
}
if (legend){
legend(legend_pos, legend=trait, col=cols, pch=pch, pt.cex=cex, bty="n", cex=legend_cex)
}
invisible(dat)
}example for multi trait
set.seed(20260108)
N <- 50000
# 1) 纯零假设(均匀分布)
p_null <- runif(N)
# 2) “多基因/少量真实信号”:97% null + 3% 偏小p
p_poly <- runif(N)
sig_poly <- rbinom(N, 1, 0.03) == 1
p_poly[sig_poly] <- rbeta(sum(sig_poly), shape1 = 0.4, shape2 = 1) # shape1<1 会产生更多小p
# 3) “膨胀”:|Z|整体偏大(sd>1),p会更偏小但不一定有真实峰
z_inf <- rnorm(N, mean = 0, sd = 1.2)
p_infl <- 2 * pnorm(-abs(z_inf))
# 4) “强信号”:1% 极强关联(生成极小p),并故意加入几个0
p_strong <- runif(N)
idx_strong <- sample.int(N, size = round(0.01 * N))
p_strong[idx_strong] <- 10^(-runif(length(idx_strong), min = 6, max = 10)) # 1e-6 到 1e-30
# p_strong[sample.int(N, 5)] <- 0 # 故意放几个0,测试防 Inf
pvals_list <- list(
Null = p_null,
Polygenic = p_poly,
Inflated = p_infl,
Strong = p_strong
)绘图
png("qqplot_multi.png", width=2400, height=2400, res=500, type="cairo")
qqplot_multi(
pvals_list,
cols = c("#1b9e77", "#d95f02", "#7570b3", "#e7298a"),
pch = c(16, 17, 15, 18),
cex = c(0.8, 0.8, 0.8, 0.8),
ci_scale = seq(0.8, 1.0, length.out = 4),
ci_alpha=0.6)
dev.off()